 December
03
December
03
Tags
Meet Christopher Lee – Shape matters! Stories of a “model” scientist
What does Lotso, the bear from Pixar’s Toy Story 3, have in common with the research of computational scientist Dr. Christopher Lee? Lotso is created with a similar technology that Christopher uses to create real 3D models of neurons.
For those who just started following this interview series, I am a researcher myself and conduct interviews with fellow scientists and then tell you about their research and their stories as scientists.

This time, I talked to Christopher Lee, a biochemist and computational scientist in the Department of Mechanical and Aerospace Engineering at UC San Diego. How Christopher became a computer scientist and arrived at this department to work on neuroscience wasn’t straightforward. At the beginning of his university career research wasn’t Christopher’s plan. Initially, he wanted to become a medical doctor, because it was the kind of “popular” thing to do. He took all the classes and bought all the expensive books to prepare for the entry exam to medical school, but at the end of his undergrad, he realized that going to medical school and learning a bazillion of medical terms, wasn’t what he wanted. Instead he was drawn to do research but didn’t exactly know how to get into this career path.
Thanks to some advisers at university that showed him that research is a real career possibility, he took a class called “Teach Science as you would do Science,” where students worked on real scientific problems, like how the structure of a protein would relate to its function (1). A cell produces millions of proteins and it’s their functions that make life possible. Their functions are often dictated by their shape, like the shape of a spoon makes it the perfect tool to eat soup. By comparing the shape of proteins of unknown function to known examples, researchers may decipher what they actually do in the cell. The use of computer programs for this type of work didn’t really scare Christopher. “Earlier on I developed some interest in writing [computer] programs, because I found that you can solve problems much easier using software,” he tells me and adds: “One could be really creative and there is an elegance of the types of ideas that one can come up to solve these problems and that really inspired me and I thought – this is actually meaningful and I could do this as a career.” So, after completing his Bachelor’s degree, he started his PhD studies in chemistry, where he worked on creating computer models of membrane structures that enclose cells (2).
As scientists, we use models all the time. Models are representations of real natural structures, like a globe. They help us to simplify complicated natural structures or allow us to visualize things that are normally invisible to the naked eye. We make models from whole organisms, cells, and even the tiniest molecular structures like proteins and membranes, such as Christopher did. Many of these models only show us a gross overview of the structure we look at; other models give us information down to the atomic level. But models used in biology are not just static like a globe, computer models can be very dynamic. The beauty of those computer simulations is that they allow researchers to test ideas or hypotheses without having to do experiments in the laboratory that are often very costly and time consuming. Later, computer simulations guide how scientists plan experiments in a laboratory (3). This is particularly important for drug discovery. Take the Covid-19 virus as an example: We know how it looks and how it infects cells. Scientists also know the structure of many drugs that have already been used before. What computer models can do is checking if a drug interrupts the contact of the virus with the host cell, just based on its shape and the virus’ shape. This gives the chance to find many potential drug candidates (4).
Christopher started with modeling parts of cells, but in the end of his PhD he got more and more interested in larger biological systems, like cells. He hypothesized that the shape of a cell matters for the function of the cell. Think about it – it makes a difference if a signal has to bridge the distance of a long-shaped or a round-shaped cell.
He went to a meeting with a professor across campus to talk about his ideas. Little did he know that this was going to be his future research mentor who was working on this type of concept. That shape matters might be particularly important in the brain, which is transporting signals all the time. Christopher is pursuing the idea of how we learn. “It turns out that neurons () have very unique geometries. () The thing is, we still don’t really understand how the properties of the neurons necessarily give rise to cognition and intelligence and things like that.” That’s what models of neurons could help answer.
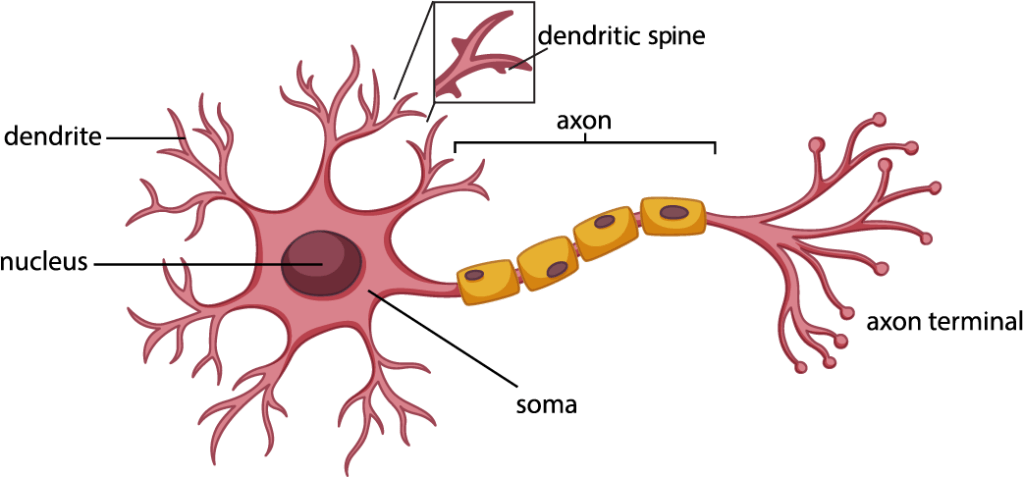
Christopher creates scientific models of very specific structures of neurons, the so-called dendritic spines. When you look at models of neurons, they look a bit like tiny trees (see Figure 1). The control center of neurons is the cell body, or soma, where signals are received from other neurons through the branches, the dendrites. Dendrites can have little protrusions, the dendritic spines, which work almost like little antennas to receive signals from neighboring neurons. Spines are extremely important to build up memories and are therefore very dynamic structures. They can vary in size, shape and ability to receive signals (5). Think of a real tree: if you place this tree in different environments – like a very shady place versus a sunny one – the same tree might sprout different amounts of branches. Neurons are also influenced by many factors that change their shape and size. This is where Christopher comes in – he wants to be able to predict in a computer simulated model which shape dendritic spines are going to take in response to certain signals.
Christopher bases his models on very detailed microscopic pictures of this small area of neurons. The method that is used to make these pictures is called “focused ion beam scanning electron microscopy”. What sounds like something out of a science fiction movie is a really cool technique that has been developed in the last decade. The sample you want to look at gets encased in a block of a polymer and with a very focused ion beam, one layer at a time of the sample gets etched away. Every time one layer is gone, the microscope takes a picture and then etches the next layer and so on and so forth. We are speaking here of the tiniest of steps down to a nanometer scale, which is a million times smaller than a millimeter. “If you want the full 3D of one cell, you need to go from top to bottom and depending how thin your slices are that could be thousands of layers just for one cell,” Christopher explains. “Those pictures need gigabytes to terabytes of space on a hard disc.”
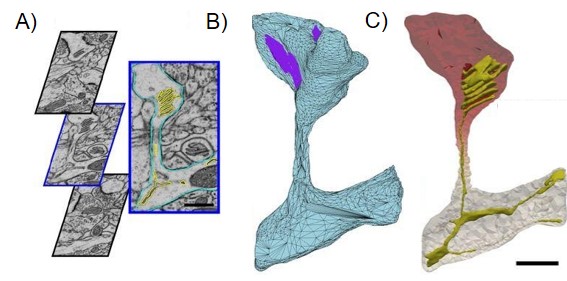
Using these series of snapshots, he starts his computational modeling. The problem here is the vast amount of data and also the difficulties to distinguish cells from the background. For untrained people all these pictures might more or less look like different shapes in different shades of grey (see in A in Figure 2). “Often you can use computer algorithms to predict where these boundaries (of cells) are, but that doesn’t always work that well, so we have to manually create those things,” Christopher adds.
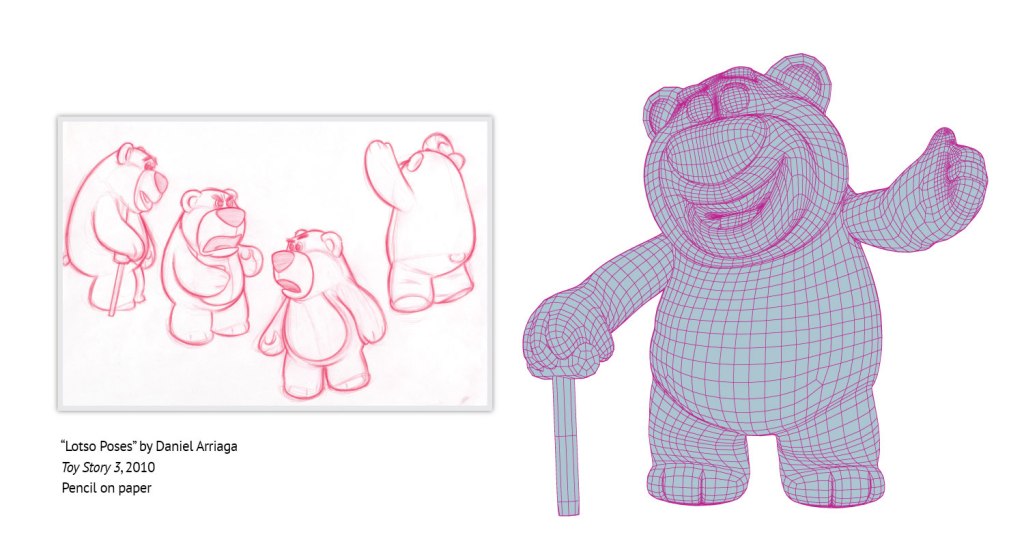
How does Christopher create surface models from these pictures? “When you go and watch these Pixar movies and you see a character’s head or hand – all this skin and those surfaces are presented as vertices with edges that connect them. You have a connected surface which we would call a surface mesh,” Christopher explains. You can see an example of such a surface mesh in the picture below of Lotso, the bear, from Pixar (Figure 3), which is later filled to create a 3D computer animation. Christopher uses this technology to model the surface of the dendritic spines and also structures inside the cells (Figure 2B and D) which gives a very realistic 3D representation of the actual shape (Figure 4).

What could these models be used for? These realistic models could be used to test memory processes in the neuron and that again could inspire drug discovery to influence neurons and thereby the function of the brain. But computer simulations that model the activity of the brain might also revolutionize computer systems themselves. “The brain is an amazingly low energy organ, compared to running a GPU (graphics processing unit – used in a computer to process images) which is running really hot, especially right now when I am running simulations at home. So, maybe we could use this (technology) to design new computational devices.” And it is true, even though the brain consumes a lot of the energy in our body, it still functions way more efficiently than our best computers. Software developers already use the principles of neuronal networks for building “intelligent” computer programs. A more detailed understanding of those might allow scientists and engineers to increase the efficiency of computers and other technology.
In some people’s minds, computer scientists are solitary nerds, sitting every day in front of their computers and writing codes on a black screen. Christopher completely dispels this picture of computer scientists (even though his daily work certainly involves a lot of computer work and coding). When I asked him, what excites him most about research, he promptly answered: “The thing to work with other smart people. There are so many interesting contributions from all folks on all levels.” This doesn’t only involve other scientists. “We have a lot of efforts in outreach even on high school level and these high school students come in and teach me new things that I didn’t think about,” Christopher adds. This stimulating environment is particularly evident in his current laboratory led by Dr. Padmini Rangamani, where chemists, physicists and biologists work alongside engineers and computer scientists. This is something that he wants to continue doing in his own lab in the future. Christopher loves this aspect of science – to build your own little family of people that work with you. And to say it in his own words: “It’s exciting to tackle something interesting together, right?” And I wish Christopher a “model” career as a scientist!
References:
- C Gray et al., Known structure, unknown function: An inquiry-based undergraduate biochemistry laboratory course Biochem Mol Biol Educ (2015). PMID: 26148241, DOI: 10.1002/bmb.20873
- CT Lee et al., Simulation-Based Approaches for Determining Membrane Permeability of Small Compounds, J Chem Inf Model (2016). PMID: 27043429, DOI: 10.1021/acs.jcim.6b00022
- SB Hofer et al. Dendritic spines: the stuff that memories are made of? Curr Biol (2010), PMID: 20178760, DOI: 10.1016/j.cub.2009.12.040
- CT Lee et al. Exascale Computing: A New Dawn for Computational Biology, Comput Sci Eng (2018), PMID: 30983889, DOI: 10.1109/MCSE.2018.05329812
- Y Zhou et al. Artificial intelligence in COVID-19 drug repurposing, Lancet Digit Health (2020), PMID: 32984792, DOI: 10.1016/S2589-7500(20)30192-8
- CT Lee et al., 3D mesh processing using GAMer 2 to enable reaction-diffusion simulations in realistic cellular geometries, PLoS Comput Biol (2020). PMID: 32251448, DOI: 10.1371/journal.pcbi.1007756


Pingback: Nanotechnology News – December 2020 | cultocracy