 September
10
September
10
Tags
Autism Spectrum Disorders; is all about genetics
The basics of ASD genetics
Autism Spectrum Disorders (ASD) are a group of neurodevelopmental disorders characterized by repetitive behaviors, language and communication deficits, and restricted interests. ASD has a prevalence of 2% of the population, and is usually diagnosed before age 3. One important aspect of ASD is that it shares a wide number of comorbidities with other disorders, the most common being anxiety, ADHD or depression, among many others 1. In this regard, there is an enormous diversity in the symptoms patients can show, and it is common to refer to ASD patients as ‘not one equal to another’. Indeed, this is mirrored in the same way by a great genetic heterogeneity identified in ASD cases, with no single genetic variant explaining more than 1% of the cases at best 2.
It has been suggested for a while that ASD is a genetic disorder, based on the observation by geneticists in the 90’s that twins had increased chances of sharing an ASD diagnosis compared to the general population. The clearest cases are those of monozygotic twins – the case when siblings are developed in the womb from the same zygote – which had an estimated 70% chance of sharing the ASD diagnosis 3, 4. But it wasn’t until the advent of technology that allows scientists to determine the entire set of genetic information in a person’s cells, that these postulates could be fully validated. Today, there is a great volume of research from the past two decades that clearly demonstrates that ASD is a genetic disorder that starts during early stages of human development, in the womb.
Rare and common genetic variation
In order to understand the complexity of ASD genetic causes, it’s important to know the differences between common and rare variation in genetics. Common variants are defined as polymorphisms – which is another way to denominate genetic variation – found in more than 1% of the population. These mutations in the DNA have usually mild or no effect at all in the functions of the genes in which they are located, as would be expected for variants that are shared by larger amounts of the population. Rare variants, however, are found in less than 1% of the population and tend to have some outward effect on health, and normally confer risk to particular diseases depending on their position in the genome.
The identification of rare variants is easier with current research technologies and the fact that there is an increase in the availability in the numbers of people to make comparisons and look for anomalies. Most of the efforts in the field have been to identify and characterize the genes impacted by ASD risk variants. The most common type of study to identify these rare polymorphisms in patients are called family studies, and revolve around sequencing the full genome of families with kids diagnosed with ASD, in order to be able to compare the genomes of the affected kids with the usually unaffected parents. These studies increase their ability to identify genetic differences when unaffected siblings to the ASD patient can also be added to the study, as they share a more similar background than the parents.
Rare variation it’s been widely studied in the past decades
To this day, there are a total of 102 genes identified with great confidence to be risk factors for ASD 5. But mutations in single genes don’t tell the whole story. There are other kinds of mutations that involve bigger regions of the chromosomes, called Copy Number Variants. These mutations are present in the form of deletions or duplications of big chunks of the genome, for regions that normally encompass several, sometimes dozens, of genes. As expected, this kind of mutation confers a bigger risk for several disorders, including ASD. It is now demonstrated that rare CNVs confer a substantial risk for ASD, and it has been shown in different studies that ASD patients carry an increased number of CNVs in coding regions of the genome compared to healthy subjects 6, 7.
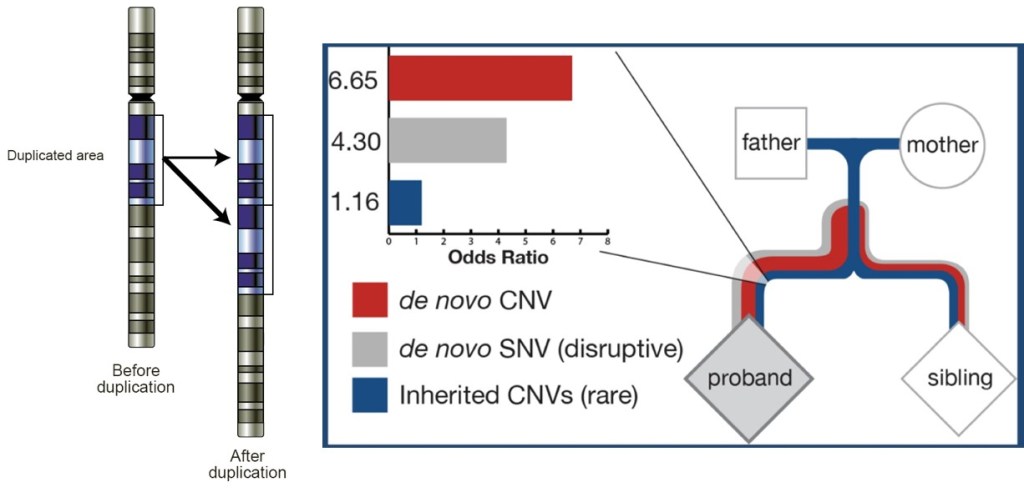
One important thing to consider is that genetic disorders are not always simply transferred from parents to children.This is a common misconception. A genetic disorder means that is caused by genetic defects, and while is true that is potentially transferable to offspring, this is not necessarily always the case. During conception, a number of variations occur in the DNA of the child that makes it different than just a mixture of 50% father and 50% mother DNA. During this process, rare pathogenic variants can appear. These are called de novo mutations, as they are formed in the new generation of offspring, without being present at all in the parents. This is what happens in most cases with ASD. It has been demonstrated now that ASD patients are carriers of more de novo deleterious mutations in genes than healthy subjects 8. This increased burden of de novo mutations in ASD patients, including both single variants and CNVs, causes disturbances in the normal development of the brain that gives rise to the symptoms observed in ASD patients later in life.
ASD risk genes can be grouped depending on their role
Given the large number of different genes identified to date that confer risk for ASD, and likewise, the large number of CNVs associated with related disorders, the next big question is how to make sense of the biology associated with this neurodevelopmental disorder, considering there are so many players involved. How is it possible that such a vast number of mutations on different genes on totally different people, can yield similar traits?
Luckily for the understanding of our limited brains, and also for researchers in the field, it happens that most of the genes affected by ASD mutations converge in their functions 9, 10. Indeed, the genes identified can be grouped depending on their role in health and brain function. Many of the risk genes associated with ASD to date fall under two categories: associated with neuronal functions, ranging from regulation of neuron formation and its network assembly. or associated with gene transcription and regulation of DNA structure. One example of the former group would be the gene SCN2A, a voltage-gated sodium channel responsible for the regulation of the transmission of information between neurons. An example of the latter group would be the gene CHD8, which regulates the transcription of several genes. Of course this is not set in stone, and there are a lot of ‘fringe’ risk genes that escape this simplistic definition or have other functions. But it’s a good way to highlight the importance of proper regulation of all the processes regulating development, and how dysregulation of any of the many genes involved can have pathogenic consequences.
Common variation in ASD
Finally, it’s important to not underestimate the relevance of common genetic variations in ASD genetic etiology. While most of the focus of research has been on the identification of rare variants, it has long been accepted that common variations are an important part of ASD risk 11. However, the studies that can identify loci of risk for common variation, require a big sample size that until recently has not been possible to accomplish for ASD studies. It wasn’t until last year that the first successful study identifying common variants was published 12. In total, five different loci for common variation were identified to confer risk for ASD, and they were located close to genes related to the functions previously mentioned, further supporting those conclusions. Most interestingly, it was also found that these loci were shared between schizophrenia and depression disorders, something that was long hypothesized given the high degree of comorbidity between these disorders and ASD.
How do common and rare genetic variations interact in the context of ASD risk? It is clear that rare risk ASD variants are not 100% penetrant. This means that not everyone with one, or even several, of these variations in their genome will develop the disorder. This is because common variation in our genome confers a genetic background that will confer more or less vulnerability for these rare mutations. In summary, while the presence of risk variants in our genomes is a defining factor on whether we will develop ASD, it is the common variation in the entire genome that dictates the extent of the effect this rare variant can have over development.
Despite the great deal of knowledge gained in the last years about ASD genetic variants and their effects, there are still many questions unanswered and many more risk genes are likely to be discovered. We hope the efforts of the research community help to shed light in this regard, with the hope to improve the lives of the patients affected by ASD, and to dispel misinformation and myths regarding these disorders.
References
1. Stavropoulos, K.K., Bolourian, Y. & Blacher, J. Differential Diagnosis of Autism Spectrum Disorder and Post Traumatic Stress Disorder: Two Clinical Cases. Journal of clinical medicine 7 (2018).
2. Geschwind, D.H. & State, M.W. Gene hunting in autism spectrum disorder: on the path to precision medicine. The Lancet. Neurology 14, 1109-1120 (2015).
3. Bailey, A. et al. Autism as a strongly genetic disorder: evidence from a British twin study. Psychological medicine 25, 63-77 (1995).
4. Bolton, P. et al. A case-control family history study of autism. Journal of child psychology and psychiatry, and allied disciplines 35, 877-900 (1994).
5. Satterstrom, F.K. et al. Large-Scale Exome Sequencing Study Implicates Both Developmental and Functional Changes in the Neurobiology of Autism. Cell 180, 568-584.e523 (2020).
6. Sebat, J. et al. Strong association of de novo copy number mutations with autism. Science 316, 445-449 (2007).
7. Sanders, S.J. et al. Multiple recurrent de novo CNVs, including duplications of the 7q11.23 Williams syndrome region, are strongly associated with autism. Neuron 70, 863-885 (2011).
8. Sanders, S.J. et al. Insights into Autism Spectrum Disorder Genomic Architecture and Biology from 71 Risk Loci. Neuron 87, 1215-1233 (2015).
9. Quesnel-Vallières, M., Weatheritt, R.J., Cordes, S.P. & Blencowe, B.J. Autism spectrum disorder: insights into convergent mechanisms from transcriptomics. Nat Rev Genet 20, 51-63 (2019).
10. Pinto, D. et al. Convergence of genes and cellular pathways dysregulated in autism spectrum disorders. Am J Hum Genet 94, 677-694 (2014).
11. Gaugler, T. et al. Most genetic risk for autism resides with common variation. Nat Genet 46, 881-885 (2014).
12. Grove, J. et al. Identification of common genetic risk variants for autism spectrum disorder. Nat Genet 51, 431-444 (2019).
13. Krumm, N. et al. Transmission disequilibrium of small CNVs in simplex autism. Am J Hum Genet 93, 595-606 (2013).

Pingback: Nyheder2020september10 - bionyt.dk