 February
21
February
21
Tags
The Scientific Adventures of Ben Barres Part I: Dishing out Discoveries
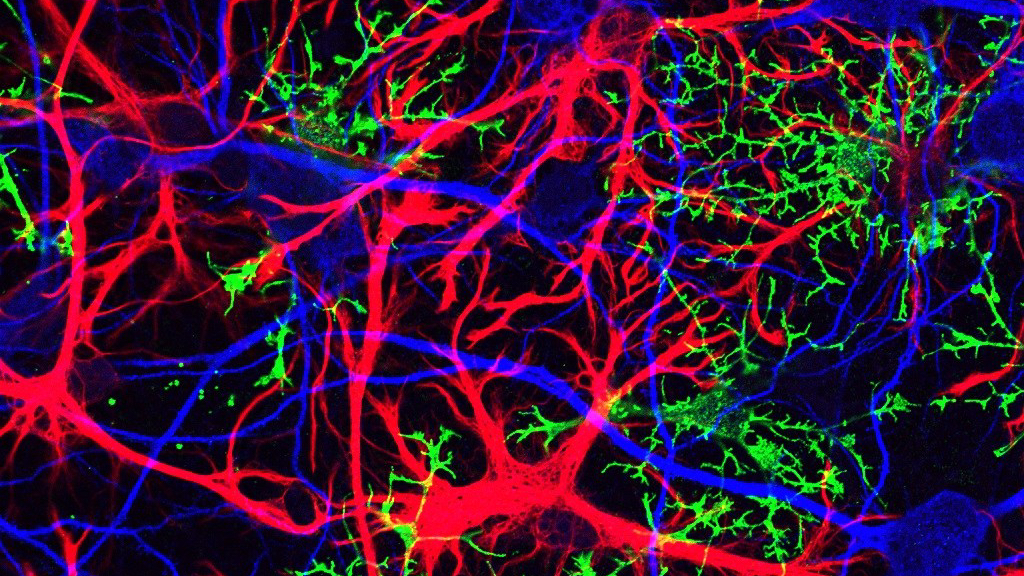
Dr. Ben Barres is a hero to those of us here at NeuWriteSD. Ben was a pioneering scientist and a devoted mentor, as well as a relentless crusader for greater inclusivity in science. While we’ve highlighted his advocacy work and published a memorial post after his death, we haven’t paid nearly enough tribute to his magnificent science career, or to the glial cells he found so enticing. The term “glia” refers to the several cell types in the brain and spinal cord that are not neurons. What the field now knows about the vital role of glial cells is due in large part to the work of the Barres lab.

Published posthumously, The Autobiography of a Transgender Scientist details Ben’s early life and role in advocacy but also takes its reader along for a wild journey of scientific discovery. In the book’s introduction, Ben writes, “[…] I would like to tell you about glial cells, what we know as well as what we don’t know, and the great adventure it has been to explore their roles in healthy and diseased brains.” The story of Ben’s science, however, is written for an audience familiar with the unique shorthand of a cellular and molecular biologist. I hope to make it a bit more accessible as I take you along on the adventure of Ben’s stellar career in a two-part series highlighting the Barres lab’s accomplishments.
Before we begin, I must emphasize Ben’s humility in the face of incredible scientific success. While it is customary for the head of a lab to take credit for what his trainees discover, Ben never did so. This was true in life and also in his autobiography; throughout the science section of his book, Ben is careful to name the graduate student or postdoctoral fellow in his lab who made each discovery. As I know this was important to Ben’s philosophy of mentorship, I will follow his example by doing the same.
The Barres Lab: A Culture of Culturing
The term “cell culture” simply refers to keeping cells alive in a dish. This may sound straightforward, but often it is no small feat. The dish must be kept in a warm incubator with a precise ratio of oxygen, nitrogen, and carbon dioxide, mimicking the body environment in which the cells thrive. Another vital aspect of keeping cells alive and happy is their growth media—the liquid that covers them in the dish. Some cell types are very picky and will die within hours without an extremely specific cocktail of molecules in their media bath.

Working with cultured cells is a central aspect of many branches of biology and pharmacology, including neuroscience. Understanding how cells behave in the body often first involves manipulating them in culture. Scientists can add drugs to block specific cellular pathways, insert a particular gene into the cells, add inflammatory signals or support signals to the cells—the list goes on and on. After any of these manipulations, the researchers can observe the cells under a microscope, looking for changes in their behavior, their growth rate, the strength of their connections to one another, etc. All of this is quite easy to do in culture, and extraordinarily difficult to do in a living animal!
At the time Ben started his own lab at Stanford in 1993, no one had been able to get central nervous system (CNS) neurons—neurons in the brain, spinal cord, and retina—to grow in a dish, despite trying endless media cocktails. CNS neurons could survive when they were accompanied by lots of other glial cell types that surrounded them in the living brain, but alone they would die almost immediately. One of Ben’s early trainees, Anke Meyer-Franke, was the first to crack the code. She combined several growth factors (common media components—usually steroids—that induce cell growth) with a drug that stimulated electrical activity in the neurons. Suddenly the CNS neurons from the rat survived for weeks in a dish, happily extending beautiful axons and dendrites, the thin extensions of their bodies that allow them to interact with one another [1]. Extending this finding, an MD/PhD student in the lab, Jeff Goldberg, showed that the growth of CNS neurons was increased almost ten-fold when the cells were grown on silicon chips that simulated the levels of electrical activity in the brain [2]. Together, Anke’s and Jeff’s discoveries suggest that CNS neurons need to stay active in order to stay healthy!
Because of the tenacity of some of Ben’s earliest lab members, the world could now, for the first time, take neurons from the rodent CNS and get them to survive and grow in a dish by themselves. This would allow neuroscientists to finally manipulate brain, spinal cord, and retinal neurons in order to discover more about how they work, how they interact with one another, and what can hurt them. Importantly, it would also allow scientists—including future Barres lab members—to add different glial cell types to the neurons one-by-one and thus determine exactly what effects glial cells had on neurons.
A Switch at Birth
One of the first new discoveries to come from studying CNS cells in a dish was that CNS neurons taken from rats of different ages behave quite differently! Jeff found that while neurons from a rat embryo extend axons (the part of the neuron that sends signals to its partners) in culture, cells purified from rats at the time of birth or later largely lose this ability. Instead, they primarily extend dendrites (the part of the neuron that listens to signals from other cells) [3].
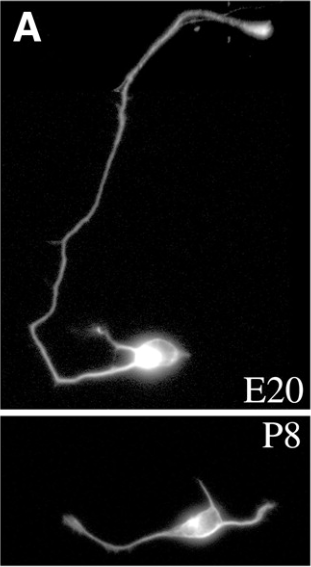
This has important implications for nerve regeneration after injury—it is likely that CNS injury is so devastating because neurons lose an intrinsic ability extend their axons. For instance, a spinal cord injury often results in paralysis because the injury severs axons of brain neurons sending signals to the limbs to move, and those axons do not regenerate. Before Jeff’s discovery, most scientists believed that axon regeneration was blocked by tissue scarring, but Jeff’s work suggested that it might be a natural, developmental change in the neurons. Many labs are now hard at work trying to understand how to reprogram these axons to re-grow.
A Great Goldmine of Data
The feat of coaxing CNS cells to survive in culture was not the only important technique that Ben’s lab pioneered. Ben was particularly interested in astrocytes, a type of glia whose seemingly endless roles in health and disease will be discussed in Part II of this post. However, no one had ever been able to obtain a pure cell culture of just astrocytes from the mix of neurons and other glial cells in the rodent brain. Ben’s trainee John Cahoy figured out how to do it. With a couple new techniques up their sleeves, John and Ben Emery not only got the purified astrocytes they were after, but ended up with separate, pure populations of neurons and oligodendrocytes (another type of glia) as well. They were then able to generate a dataset of all of the genes expressed by each of these cell types during the development of the rat [4]. Despite all of the magnificent discoveries to come out of the Barres lab, the paper detailing these data is still the most heavily-cited of them all, meaning that many other scientists have used this knowledge as a basis for their own discoveries.
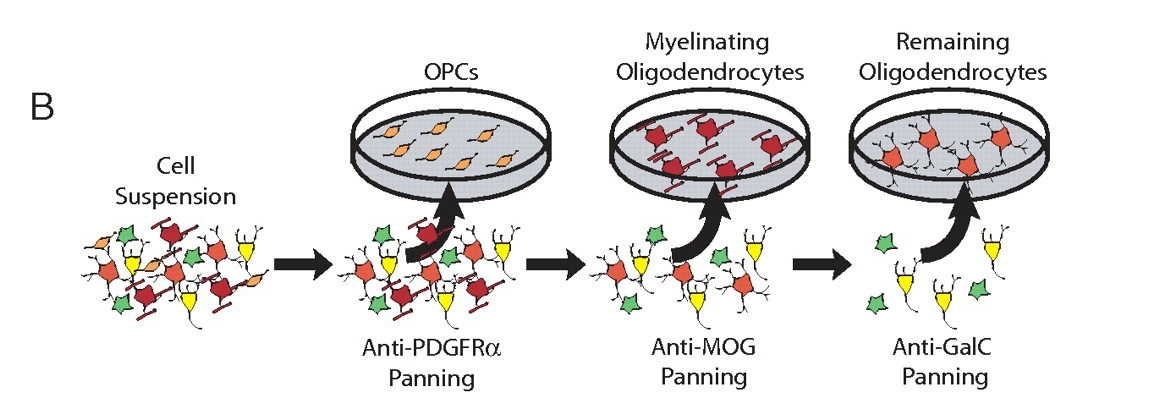
Scientific results are often judged by the extent to which they “move the field forward.” This was a huge leap for the field of neuroscience. A few other cell types were later added to the database, and the entire thing was made easily accessible to the public on a webpage. Anyone can search any gene of interest and the site will immediately spit out a graph showing which cell types express the gene during brain development. Its instantaneous results are addicting… I may or may not have the shortcut to this webpage on the homepage of my phone.
My Oh Myelin
A significant portion of the Barres lab studied oligodendrocytes, glial cells that create myelin and ensheath neurons with it. Myelin is a fatty substance that insulates the neurons, allowing them to conduct current far more quickly. Without myelin, neurons cannot signal to one another effectively. Indeed, the devastating physical and cognitive symptoms of multiple sclerosis (MS) arise from myelin damage.
As a graduate student in the Barres lab, Trent Watkins discovered a particular compound that could rapidly induce myelination in cell culture, a process that he could watch with time-lapse imaging [5]. Surprisingly, he found that once an oligodendrocyte decides to produce myelin, it myelinates all of its target neurons within a few days, and then never produces myelin again. This is vital information as it suggests that re-myelination for MS patients will require generating new oligodendrocytes altogether, because mature oligodendrocytes are not able to re-myelinate.
Several other findings from the Barres lab have added to what we know about myelination. Jason Dugas found that an oligodendrocyte goes through many distinct stages before it can start myelinating [6], and Ben Emery discovered a particular oligodendrocyte gene required for myelination [7]. These and other discoveries provide key information about oligodendrocytes that will hopefully aid in therapeutic strategies for re-myelination in MS patients.
Blowing Open the Blood-Brain Barrier Field
Another cell type that begs recognition is the endothelial cell. Endothelial cells make up the walls of the blood vessels that bring oxygen and nutrients to every nook and cranny of the body, including the brain. However, the blood vessels in the brain are special—unlike other endothelial cells, brain endothelial cells form blood vessels with an especially tight seal, controlling what can move from the blood into the main tissue of the brain. This tight control is called the blood-brain barrier (BBB for short), star of its own NeuWriteSD post.
Ben’s lab did not discover the BBB, but one of Ben’s graduate students, Richard Daneman, exponentially expanded what the world knows about it. He isolated endothelial cells from the brain and from several other organs. He then identified the full set of genes expressed uniquely by endothelial cells in the brain [9], providing a vital resource for understanding how and why endothelial cells in the brain act so differently from those in the rest of the body.
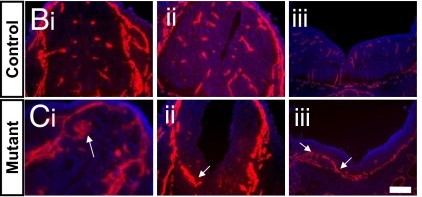
Rich also made several discoveries about how the BBB forms during development. Researchers had long assumed that astrocytes were the cells responsible for inducing the specialized functions of brain endothelial cells that make up the BBB. However, Rich discovered that the BBB actually forms before astrocytes arrive on the scene, while mice lacking a different cell type, pericytes, never form a BBB and die before birth [10]. This turned the field upside-down, showing definitively that pericytes were indispensable for BBB formation. He also found that a particular molecular signal (Wnt) is not only essential for inducing some BBB properties, but it is also required for blood vessel formation in the CNS. This was quite surprising as Wnt is not needed for blood vessel formation in the rest of the body. Rich’s discovery adds another element to our understanding of just how unique CNS vasculature is!
Stay Tuned!
This marks the end of the first half of our adventure through Ben Barres’ incredible career of discovery. His lab figured out how to keep CNS neurons alive in a dish, allowing neuroscientists around the world to study them more closely. The lab also pushed the field of neuroscience further by generating huge datasets describing which genes are turned on by various CNS cells during development.
Next week, the second half of our adventure will take us to the stars. The star cells, that is. Are astrocytes good or evil? Stay tuned!
References
1. Meyer-Franke A, Kaplan MR, Pfrieger FW, Barres BA. Characterization of the signaling interactions that promote the survival and growth of developing retinal ganglion cells in culture. Neuron. 1995 Oct;15(4):805-19. PubMed PMID: 7576630.
2. Goldberg JL, Espinosa JS, Xu Y, Davidson N, Kovacs GT, Barres BA. Retinal ganglion cells do not extend axons by default: promotion by neurotrophic signaling and electrical activity. Neuron. 2002 Feb 28;33(5):689-702. PubMed PMID: 11879647.
3. Goldberg JL, Klassen MP, Hua Y, Barres BA. Amacrine-signaled loss of intrinsic axon growth ability by retinal ganglion cells. Science. 2002 Jun 7;296(5574):1860-4. PubMed PMID: 12052959.
4. Cahoy JD, Emery B, Kaushal A, Foo LC, Zamanian JL, Christopherson KS, Xing Y, Lubischer JL, Krieg PA, Krupenko SA, Thompson WJ, Barres BA. A transcriptome database for astrocytes, neurons, and oligodendrocytes: a new resource for understanding brain development and function. J Neurosci. 2008 Jan 2;28(1):264-78. doi: 10.1523/JNEUROSCI.4178-07.2008. PubMed PMID: 18171944.
5. Watkins TA, Emery B, Mulinyawe S, Barres BA. Distinct stages of myelination regulated by gamma-secretase and astrocytes in a rapidly myelinating CNS coculture system. Neuron. 2008 Nov 26;60(4):555-69. doi: 10.1016/j.neuron.2008.09.011. PubMed PMID: 19038214; PubMed Central PMCID:PMC2650711.
6. Dugas JC, Tai YC, Speed TP, Ngai J, Barres BA. Functional genomic analysis of oligodendrocyte differentiation. J Neurosci. 2006 Oct 25;26(43):10967-83. PubMed PMID: 17065439.
7. Emery B, Agalliu D, Cahoy JD, Watkins TA, Dugas JC, Mulinyawe SB, Ibrahim A, Ligon KL, Rowitch DH, Barres BA. Myelin gene regulatory factor is a critical transcriptional regulator required for CNS myelination. Cell. 2009 Jul 10;138(1):172-85. doi: 10.1016/j.cell.2009.04.031. PubMed PMID: 19596243; PubMed Central PMCID: PMC2757090.
8. Daneman R, Zhou L, Agalliu D, Cahoy JD, Kaushal A, Barres BA. The mouse blood-brain barrier transcriptome: a new resource for understanding the development and function of brain endothelial cells. PLoS One. 2010 Oct 29;5(10):e13741. doi: 10.1371/journal.pone. 0013741. PubMed PMID: 21060791; PubMed Central PMCID: PMC2966423.
9. Daneman R, Zhou L, Kebede AA, Barres BA. Pericytes are required for blood-brain barrier integrity during embryogenesis. Nature. 2010 Nov 25;468(7323):562-6. doi: 10.1038/nature09513. Epub 2010 Oct 13. PubMed PMID:20944625; PubMed Central PMCID: PMC3241506.
10. Daneman R, Agalliu D, Zhou L, Kuhnert F, Kuo CJ, Barres BA. Wnt/beta-catenin signaling is required for CNS, but not non-CNS, angiogenesis. Proc Natl Acad Sci U S A. 2009 Jan 13;106(2):641-6. Doi 10.1073/pnas.0805165106. Epub 2009 Jan 7. Erratum in: Proc Natl Acad Sci U S A. 2009 Apr 14;106(15):6422. PubMed PMID:19129494; PubMed Central PMCID: PMC2626756.

Enjoyed reading this Daisy!!
On Thu, Feb 21, 2019 at 8:24 PM NeuWrite San Diego wrote:
> Catie Profaci posted: ” Astrocytes (red) and oligodendrocytes (green) are > two types of glial cells that the Barres lab studied. These glial cells > have profound effects on neurons (blue). Image from Jonathan Cohen at the > NIH Dr. Ben Barres is a hero to those of us here at” >
Pingback: Your brain is plastic! | NeuWrite San Diego