 December
10
December
10
Tags
A Toast to Optogenetics
“This seems rather far-fetched but it is conceivable that molecular biologists could engineer a particular cell type to be sensitive to light.”

Happy 10-year anniversary, Optogenetics!
These words, published in 1999 by Francis Crick [1] (co-discoverer of the DNA double helix structure and a neuroscientist later in life) were incredibly prophetic. It did seem far-fetched, and yet, a mere six years after that, the first paper was published showing that mammalian neurons could be reliably activated with light. Here we are, 10 years later, and optogenetics – opto for “light” and genetics to refer to the fact that neurons are made light-sensitive by controlling gene expression – has rapidly become one of the most widely used tools in neuroscience. In honor of the 10th anniversary, let us celebrate as if optogenetics were not merely a scientific tool but rather a loved one celebrating 10 years of marriage. Let us raise our glasses for a toast and reminisce on how it all began!
But first, why the celebration? Why is optogenetics such a powerful tool that it merits an anniversary celebration at all? Imagine that you are a neuroscientist and you want to determine whether a population of neurons is important for something like memory or attention. To do that, you would need to inactivate that population and observe whether the behavior becomes impaired in order to establish that those neurons are necessary for that behavior. Depending on how strong a claim you want to make, you might even want to establish sufficiency, that activation of those neurons produces that behavior. This would require you to manipulate those neurons’ activity specifically and precisely. Manipulation has to be specific if you want to establish that those neurons alone are important for that behavior, and it has to be precise on a millisecond timescale that mirrors the activity of actual neuronal activity or else risk throwing off the entire system. Unfortunately, if you had wanted to do these experiments more than 10 years ago, you wouldn’t have been able to.

Thanks for your invention, Ed and Karl!
This brings us to the beginning of our anniversary slideshow, circa 2000, where we see Stanford students Ed Boyden and Karl Deisseroth pining over this very scenario. Many neuroscientists were trying to figure out how to specifically and precisely activate neurons, but most of the ideas involved magnetic or electric fields that were impossible to restrict to a single-neuron scale. Though only doctoral students at the time, Boyden and Deisseroth had the idea that light could be more precise, if only there was some way to make neurons light-sensitive…
The solution began to emerge when Boyden stumbled upon a whole field of literature on microbial opsins, which are proteins found in single-celled organisms, such as algae, that convert light into electrical energy. Most of the organisms in which these proteins had been found flourished in extremely salty environments, but for the first time in 1999, one particular protein was found whose optimal conditions were similar to the natural conditions inside the brain [2]. A few years later in 2003, a group in Germany published a paper on the successful use of a new light-sensitive protein found in green algae, channelrhodopsin in cultured mammalian kidney cells [3]. These findings hinted that perhaps these proteins could be viably expressed in neurons.
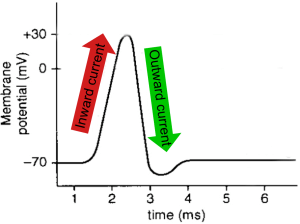
Fig. 1: The characteristic “spike” shape of an action potential is the result of an inward current of positively-charged sodium ions into the neuron followed by an outward current of positively-charged potassium ions out of the neuron.
To fully appreciate the utility of activating neurons with light and how this might be achieved, it is necessary to understand that activity in the brain takes the form of electrical signals. At the level of a single neuron, the most basic unit of activity is an action potential, or more colloquially, a “spike”. A spike occurs when channels in the neuron’s membrane open and trigger a rush of positively-charged sodium ions into the neuron. This quick inward current and the following outward current caused by a rapid evacuation of positively-charged potassium ions out of the cell together produce the characteristic “spike” shape (Fig 1). From there, this electrical signal travels down the neuron’s axon and ultimately initiates the neuron’s release of chemical signals. When these neurotransmitters bind to the next neuron, its electrical activity is altered, and the process continues.

Fig. 2: When blue light shines onto ChR2, it opens, allowing positively-charged ions to pass through the cell membrane. This “activates” the neuron by causing it to fire many action potentials.
Channelrhodopsin (ChR2) is a membrane channel similar to the channels controlling the neuronal spikes except that blue-wavelength light causes it to open, allowing any positively-charged ion to pass through. Thus, when blue light is shined onto neurons containing ChR2, sodium ions (which are positively-charged) are able to sneak in before the other channels in the neuron’s membrane have opened. This added charge inside the cell opens the floodgates for even more sodium to rush in and, voila! You’ve got active neurons firing many spikes at your command (Fig. 2).
But back in Palo Alto in 2003, this was merely an ideal scenario that had yet to be brought to fruition. Boyden and Deisseroth procured some ChR2 from the group in Germany and began perfecting their genetic delivery of the protein to neurons in a dish (for a nice description of how this works, see another optogenetics-themed NeuWrite post!). As it turns out, they were incredibly lucky that ChR2 was in many ways the perfect construct for what they wanted to achieve. A single gene was sufficient to make the fully-functioning protein, it safely expressed in mammalian neurons, and it worked on a fast enough timescale to be compatible with neuronal communication: altogether an incredible hat-trick of good fortune. In a classic case of feverish graduate student mania, Boyden went into his lab at 1am one morning in 2004 and for the first time recorded from a neuron expressing ChR2 that was easily activated by pulses of blue light. One year later, they published their seminal paper in Nature Neuroscience [4], thus consummating the holy academic sacrament that we are now celebrating 10 years later.

Yes, we are celebrating the anniversary of an academic paper.
In the last decade, optogenetics has matured into a ubiquitous, powerful, and remarkably easy-to-use tool for neuroscientists of all kinds. In the years following their big paper, Boyden and Deisseroth – freshly minted young faculty working sometimes in collaboration and sometimes in competition – expanded the optogenetics toolkit by adding proteins that, unlike ChR2, inactivate neurons when excited by particular-wavelength light. Today, there are many different genetically optimized opsins that are available to use (see many of them here), some of which can do things that even Crick may not have thought possible; for instance, there is a set of “step-function” opsins that only need a brief light pulse to turn neurons “on or off” for an extended period of time, waiting for another light pulse to set them back to normal.
But arguably the most critical advancement in the use of optogenetic tools has been in their application to subgroups of neurons. Recall our hypothetical, ideal scenario where we are able to reliably activate a specific population of neurons in order to determine their function. Optogenetics can be used to turn on or off a brain area to identify its role in behavior, but it can also be used to target specific types of cells within a single brain region. This allows you to determine these cells’ role in how that brain region processes information as well as their specific contribution to a behavior, since different cell types can play very different functional roles. An example of how this cell specificity is achieved is with mice that have been genetically modified to express a special protein called cre in only your cells of interest. A gene coding for an optogenetic protein, such as ChR2, can then be made cre-dependent so that when it is delivered to the mice, ChR2 will only be expressed in your cells of interest, thus allowing you to selectively turn them on with light. In this way, optogenetics has allowed modern neuroscientists to achieve a level of specificity that was simply impossible a decade ago. Incredibly, this cell-specific manipulation can also be achieved in vivo – in awake, behaving animals. By 2007, neuroscientists were able to express opsins in their cell types of interest in living mice, activating and inactivating them and observing their role in natural behavior.
In this video, ChR2 is expressed in the mouse’s motor cortex, the part of the brain that controls movement. Notice that when the blue light turns on, the mouse starts to run!
Beyond just being cool, the ability to turn specific cells on or off with light holds great promise for advancing our understanding of the brain. In many ways, it already has. For instance, it has elucidated some of the circuit mechanisms underlying our primary sensory systems like vision, audition, etc. as well as movement control, sleep and circadian rhythms, social behavior, and memory to name a few [5]. Remarkably, we have also learned a lot about various brain disorders at the cellular level. For example, optogenetics in awake mice was used to manipulate two opposing pathways deep in the brain that are involved in motor control, resulting in increased or suppressed Parkinson-like behavior. Strikingly, the selective activation of one pathway dramatically improved symptoms in a Parkinsons mouse model [6]. Recently at the annual Society for Neuroscience conference, there was a poster demonstrating that selective optogenetic activation of a specific type of cell in the brain’s frontal cortex abolished social deficits in a mouse model of autism [7]. Bear in mind that optogenetics is nowhere close to being used safely in humans for direct clinical interventions (although arguably the most promising application is for improving disrupted vision in the human retina [8]). Nevertheless, optogenetics is revealing much about how the brain works in a healthy state and what is disrupted in disordered states so that other interventions can be developed.
The relationship between neuroscientists and light-sensitive proteins has been a beautiful thing to behold. It’s amazing to think that 10 years have already gone by, and yet, so much has changed since they first joined forces that it simultaneously feels like a whole lifetime has passed. Who is to say what the next decade will bring? Improved and even more sophisticated opsins? Being able to simultaneously image and control large-scale networks in the brain [9]? New approaches for activating neurons of interest, such as with ultrasound? It’s an exciting time in neuroscience, certainly worthy of a celebratory toast.
To learn more, check out this New Yorker article, this Inquiring Minds interview with Ed Boyden, and Ed Boyden’s TED Talk.
References:
- Crick, F. (1999). The impact of molecular biology on neuroscience. Philosophical Transactions of the Royal Society of London B: Biological Sciences, 354(1392), 2021–2025. http://doi.org/10.1098/rstb.1999.0541
- D. Okuno et al. (1999). Chloride concentration dependency of the electrogenic activity of halorhodopsin. Biochemistry, 38, 5422-29.
- G. Nagel et al. (2003). Channelrhodopsin-2, a directly light-gated cation-selective membrane channel. PNAS, 100, 13940-45.
- Boyden, E. S., Zhang, F., Bamberg, E., Nagel, G., & Deisseroth, K. (2005). Millisecond-timescale, genetically targeted optical control of neural activity. Nat Neurosci, 8(9), 1263–1268. http://doi.org/10.1038/nn1525
- Deisseroth, K. (2015). Optogenetics: 10 years of microbial opsins in neuroscience. Nat Neurosci, 18(9), 1213–1225.
- Kravitz, A. V., Freeze, B. S., Parker, P. R. L., Kay, K., Thwin, M. T., Deisseroth, K., & Kreitzer, A. C. (2010). Regulation of parkinsonian motor behaviors by optogenetic control of basal ganglia circuitry. Nature, 466(7306), 622–626. http://doi.org/10.1038/nature09159
- Selimbeyoglu, C. K., et al. (2015). Optogenetic rescue of impaired social behavior phenotype in autism [Abstract]. Society for Neuroscience Annual Meeting. Retrieved from http://www.abstractsonline.com/Plan/SSResults.aspx
- Chow, B. Y., & Boyden, E. S. (2013). Optogenetics and Translational Medicine. Science Translational Medicine, 5(177), 177ps5–177ps5. http://doi.org/10.1126/scitranslmed.3003101
- Boyden, E. S. (2015). Optogenetics and the future of neuroscience. Nat Neurosci, 18(9), 1200–1201.
Other general references:
Adamantidis, A., Arber, S., Bains, J. S., Bamberg, E., Bonci, A., Buzsaki, G., … Wilson, R. I. (2015). Optogenetics: 10 years after ChR2 in neurons[mdash]views from the community. Nat Neurosci, 18(9), 1202–1212.
Boyden, E. S. (2011, July 1). The Birth of Optogenetics. The Scientist. Retrieved from http://www.the-scientist.com/?articles.view/articleNo/30756/title/The-Birth-of-Optogenetics/
Boyden, E. S. (2011). A history of optogenetics: the development of tools for controlling brain circuits with light. F1000 Biology Reports, 3, 11. http://doi.org/10.3410/B3-11
Image sources:
Title image: http://mcgovern.mit.edu/news/wp-content/uploads/2012/10/optosilenceLR2.png
Toast: http://blog.weddingcompass.com/wp-content/uploads/2012/11/Michael-Toasting-.jpg
Karl and Ed: http://www.photonics.com/Article.aspx?AID=57936
Figure 2: https://knowingneurons.files.wordpress.com/2013/02/chr2_470.jpg?w=610

Pingback: Dawn of the DREADD | NeuWrite San Diego
Pingback: How long-term memories are made - Cosmos Magazine
Pingback: How Neuroscience Tools Can Help Patients Regain Their Vision | NeuWrite San Diego